TidyMass Shiny (Online service)
An object-oriented, reproducible analysis framework for LC-MS data
Streamlined LC-MS Data Analysis
TidyMass provides a comprehensive, reproducible workflow for metabolomics and lipidomics data processing, from raw data to biological insights.
Key Features
Object-Oriented Framework
Structured data representation ensuring reproducibility and traceability throughout the analysis workflow.
Comprehensive Workflow
From raw data processing to statistical analysis and biological interpretation in one integrated environment.
Reproducible Research
Complete analysis provenance tracking with version control for all processing steps and parameters.
Concept & Workflow
TidyMass Conceptual Framework
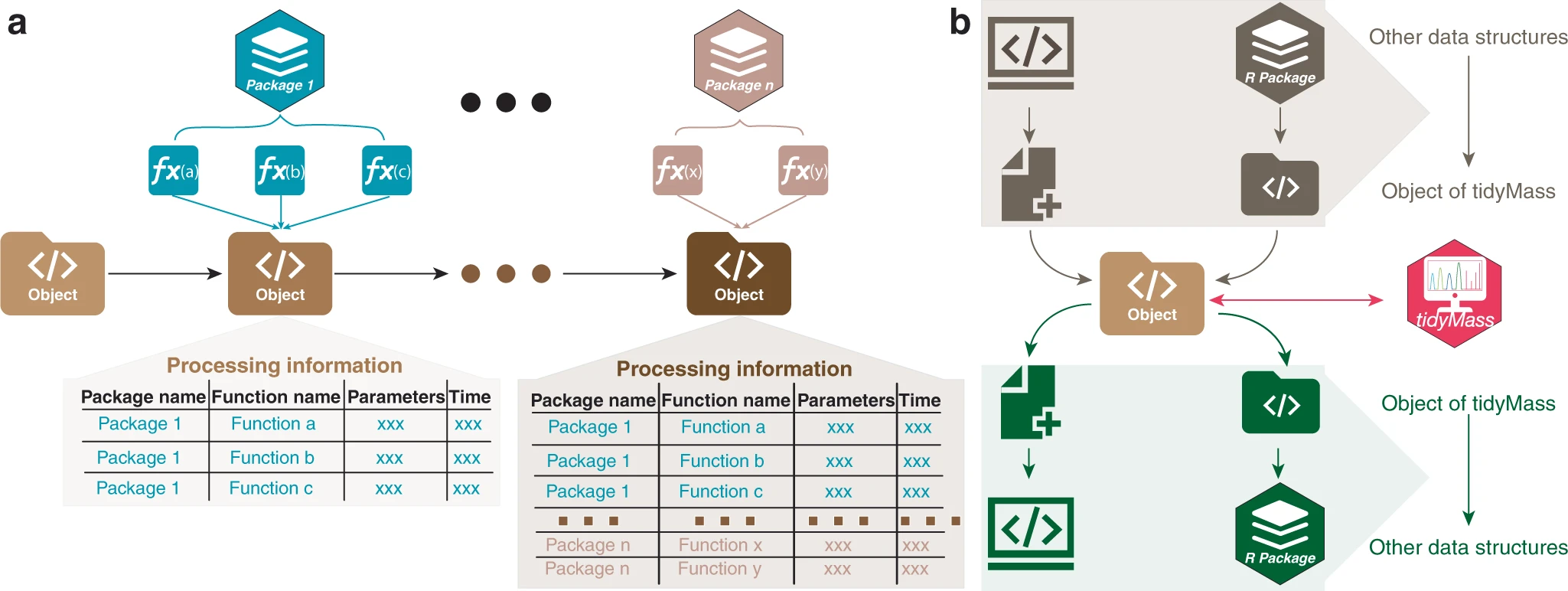
Comprehensive metabolomics data processing framework
Analysis Workflow
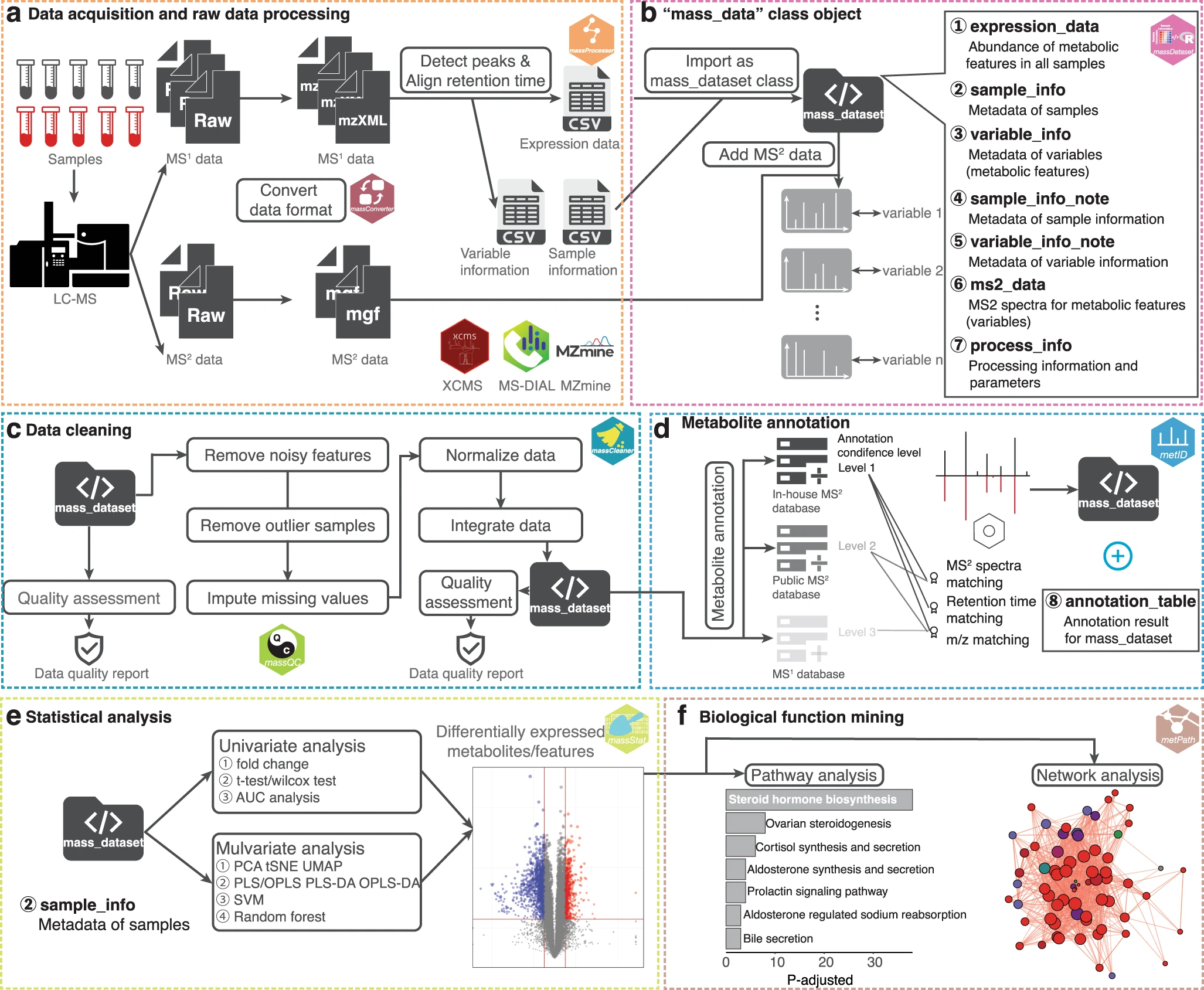
End-to-end LC-MS data processing workflow
TidyMass2 New Features
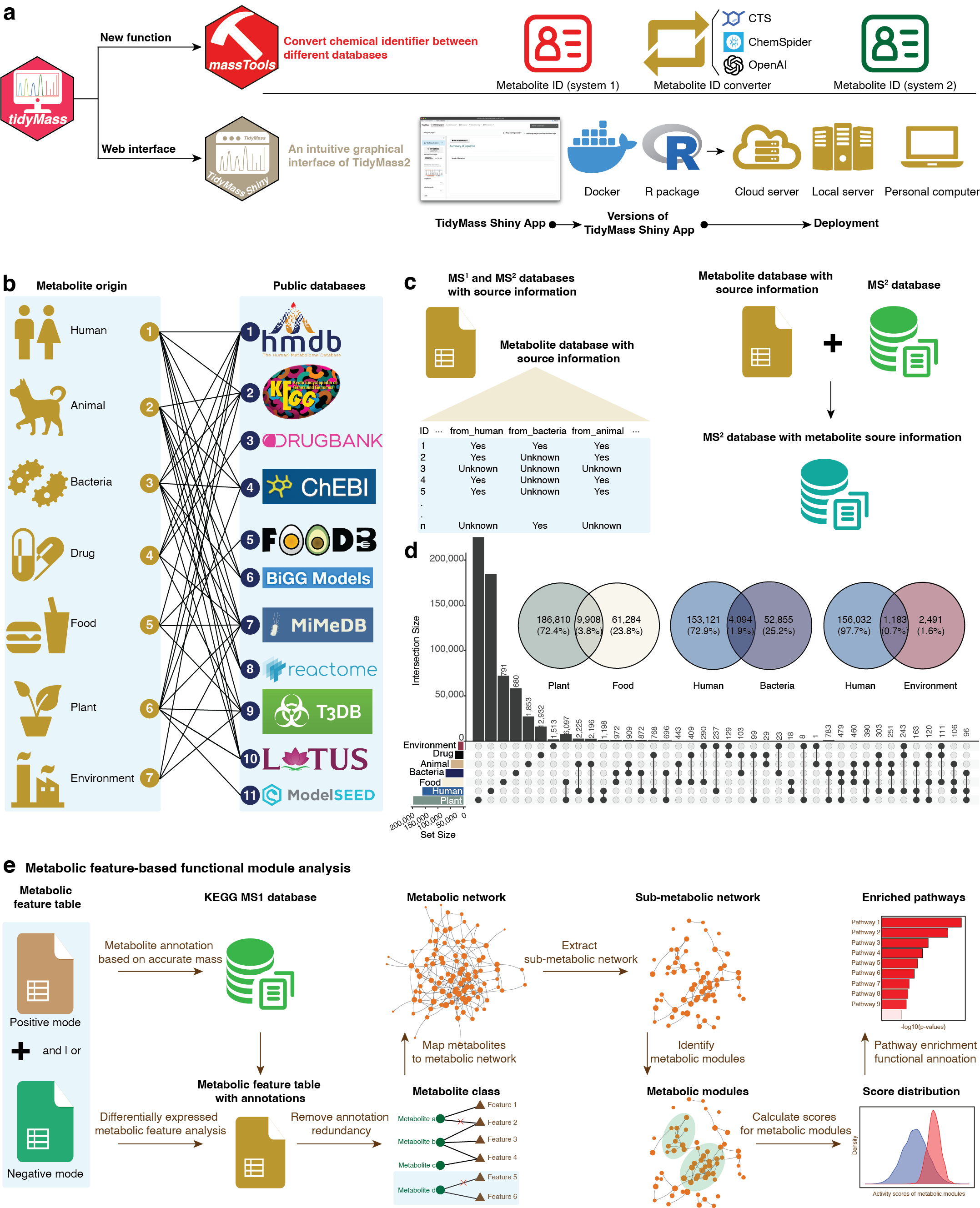
Advanced features in TidyMass2: Metabolite origin inference and metabolic feature-based functional module analysis
Key Innovations in TidyMass2:
- Cross-Platform Identifier Conversion: Comprehensive chemical identifier conversion system operating across multiple metabolite ID systems.
- Web-Based Analysis Interface: TidyMassShiny package providing a user-friendly web interface for all TidyMass2 functions.
- Metabolite Origin Inference: Integration of 11 databases into MetOriginDB for precise metabolite source categorization across 7 origin categories.
- Origin-Annotation Integration: Seamless connection between metabolite source information and MS2 spectral libraries for enhanced biological interpretation.
- Feature-Based Functional Module Analysis: Novel network approach identifying biologically relevant metabolic modules without relying solely on MS2 annotation.
- Comprehensive Metabolic Network: Human metabolic network with 9,630 metabolites and 30,196 connections for functional module detection.
Citation
If you use TidyMass in your publications, please cite:
Shen, X., Yan, H., Wang, C. et al. TidyMass an object-oriented reproducible analysis framework for LC–MS data. Nat Commun 13, 4365 (2022).
Wang, X., Liu, Y., Jiang, C. et al. TidyMass2: Advancing LC-MS Untargeted Metabolomics Through Metabolite Origin Inference and Metabolic Feature-based Functional Module Analysis.